Understanding Bacteria to Better Fight Infection

In the battle against drug-resistant bacteria, Marcos Pires studies the chemical biology of bacterial cell surfaces to better understand how they function—and possibly how to manipulate them. (Illustration by Hvass & Hannibal)
Know Thine Enemy
Alexander Fleming discovered the original antibiotic, penicillin, completely by chance in 1928. Fleming later won a Nobel Prize for his discovery, which was considered a miracle drug. However, Fleming himself recognized the potential for the development of drug-resistant bacteria.
"I would like to sound one note of warning," Fleming said during his Nobel lecture in December 1945. "... It is not difficult to make microbes resistant to penicillin in the laboratory by exposing them to concentrations not sufficient to kill them, and the same thing has occasionally happened in the body."
Fleming went on to describe a hypothetical scenario in which "the ignorant man may easily underdose himself [with penicillin] and by exposing his microbes to nonlethal quantities of the drug make them resistant." If a man with a sore throat, "Mr. X," takes an insufficient amount of penicillin to kill the streptococci bacteria causing his infection, Fleming said, he allows the bacteria to develop a resistance to penicillin. When the man's wife, "Mrs. X," becomes infected by the same bacteria, penicillin treatment will be rendered ineffective and her infection may prove fatal.
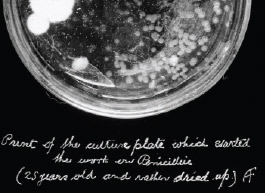
"Who is primarily responsible for Mrs. X's death?" Fleming asked. "Why Mr. X, whose negligent use of penicillin changed the nature of the microbe. Moral: If you use penicillin, use enough."
Indeed, bacteria have evolved over time, limiting the effectiveness of penicillin and many other antibiotics. In the body, these drug-resistant bacteria survive antibiotic exposure and continue to multiply. According to the Centers for Disease Control and Prevention, more than 2 million people become infected with antibiotic-resistant bacteria each year, and approximately 23,000 of those die from these infections. The World Health Organization lists antibiotic resistance as one of the biggest threats to global health today.
Marcos Pires, assistant professor of chemistry, seeks to better understand the enemy in the fight against drug-resistant bacteria. He is tackling the problem of antibiotic resistance through the creation of synthetic amino acids intended to trick bacterial cells into using some of these unnatural building blocks instead of their own. Pires believes a potential line of defense may lie in how the microorganisms construct their peptidoglycan, or cell wall. The cell wall functions like a jacket that surrounds and protects bacteria from their surroundings.
"You really have to understand how bacteria are operating, and if you do, maybe you can find a way to disrupt it," he says.
Bacterial trickery
In his initial work with bacteria, Pires and his team, inspired by the successful immunotherapy treatment of some cancers, exploited bacteria's unique use of D-amino acids to "trick" them into generating an in vitro immune response. Amino acids are the building blocks of all living organisms, and, unlike other cells, bacterial cells will incorporate D-amino acids into their cells from their external surroundings. Pires and his team created synthetic D-amino acids, tagged them with an antigen that draws a response from a pool of common human antibodies, and added the amino acids and the antibodies to a flask of bacteria. The bacteria then did the rest of the work. As they were growing, they were tricked into incorporating the lab-produced amino acids on their peptidoglycan, essentially marking themselves for destruction by the antibodies.
Pires described the results of this research in a 2014 article in The American Chemical Society's ACS Chemical Biology titled "D-Amino Acid Mediated Recruitment of Endogenous Antibodies to Bacterial Surfaces," and in a 2016 ACS Infectious Diseases cover article titled "Dipeptide-Based Metabolic Labeling of Bacterial Cells for Endogenous Antibody Recruitment."
For this work, ACS Infectious Diseases in August 2016 named Pires a winner of its Young Investigator Award for "outstanding early career researchers."
Now, encouraged by his existing work, Pires is turning to the problem of antibiotic resistance and the role of the bacterial cell wall in its development. He uses bacteria's tendency to tweak their amino acids in response to antibiotics to once again "trick" them—this time into revealing information about themselves.
Many antibiotics operate by inhibiting how bacteria put together their cell wall, says Pires. In response, bacteria modify the amino acids, or "building blocks," on their cell wall to prevent the antibiotic from attaching to it.
"That's where we come in," explains Pires. "We make that different building block [in the lab] but with a marker [to track the change.] If we make it so it's not too disruptive, [the bacteria] thinks it's picking up its own building block. But it's picking up ours."
In the same way it did in Pires' earlier research, the bacteria then places the lab-produced amino acid on its cell wall. Pires and his team add an antibiotic and monitor the process, using the marked synthetic amino acids to note any changes in the cell wall.
"That allows us to essentially follow how—or how quickly—the bacteria is becoming resistant," he explains. "We are able to see basically how long this takes."
Surprisingly, it doesn't take very long for resistance to develop. Within an hour, says Pires, bacteria adapt to escape the antibiotic. However, as soon as the antibiotic is removed, the bacteria begins returning to its original state. This demonstrates, he says, that the resistant state is "really stressful to the cell."
The team has charted this process with various types of bacteria and antibiotics to determine which types of bacteria respond to which antibiotics and how. They work with four to five types of harmless bacteria, some of which are considered "model organisms." "Usually it's the case that [when] you find something about their physiology, it will be the same more or less for the pathogenic bacteria," says Pires.
Pires believes that knowledge about bacterial physiology might help in the development of an antibiotic that can overcome changes to the cell wall. The best defense, after all, is a good offense—and a clear understanding of the enemy.
"I think now that things are a little bit more dicey [with drug resistance] and we really haven't been able to come up with new antibiotics, there's almost a revival in this [interest in] how cells are putting together their cell walls. Maybe from this we can learn how to design a drug," he says.
This story appears as "Know Thine Enemy" in the 2017 Lehigh Research Review.
Posted on:




